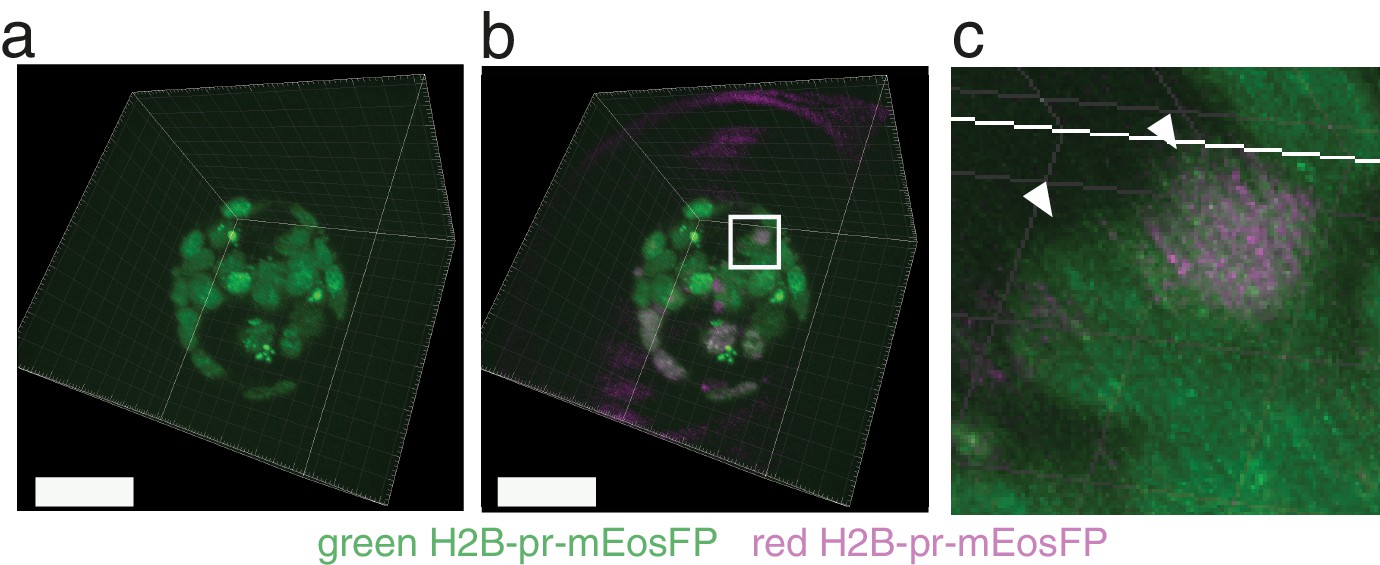
Likewise, fluorescent protein fusions that localize to the plasma membrane provide information on membrane dynamics and cell morphology. For example, fluorescent proteins fused to a human histone H2B, since they are bound to active chromatin, allow the visualization and tracking of individual cells within a group. Whereas native fluorescent proteins are cytosolic, subcellularly-localized fluorescent proteins can be used for higher resolution image information. The advent of genetically-encoded fluorescent proteins has afforded the ability to label cells ubiquitously or selectively depending on the cis-regulatory elements used in transgene design. Therefore, it has been a challenge to tag single or groups of cells that are not superficially located. Unfortunately, most of these techniques are invasive, and only effective when the tissue or cells of interest are easy accessible for manipulation. 
Several approaches have been established for labeling cells in developing mouse embryos, the mammalian genetically-tractable model of choice, including dye injections, electroporation of nucleic acids or proteins into single or groups of cells as well as grafting of genetically-distinct tissues or using chimeras. One way to gain insight into these events is to label and observe single or groups of cells over time. Genetically-encoded photomodulatable proteins such as KikGR represent emergent attractive alternatives to commonly used vital dyes, tissue grafts and genetic methods for investigating dynamic behaviors of individual cells, collective cell dynamics and fate mapping applications.Ĭell fate, pattern formation and morphogenesis depend on dynamic cell interactions involving a multitude of cell behaviors and cell populations. 
They represent novel tools for non-invasive selective labeling specific cell populations and live imaging cell dynamics and cell fate. Transgenic ES cells and mice exhibiting widespread readily detectable expression of KikGR are indistinguishable from their wild type counterparts and are amenable to efficient photoconversion. Red fluorescent photoconverted cells and their progeny could be followed for extended periods of time. By efficient photoconversion of KikGR we labeled subpopulations of ES cells in culture, and groups of cells within ex utero cultured mouse embryos. We have generated transgenic ES cell lines and strains of mice exhibiting robust widespread expression of KikGR. Our results suggest that the green-to-red photoconvertible fluorescent protein, Kikume Green-Red (KikGR), is most suitable for cell labeling and lineage studies in ES cells and mice because it is developmentally neutral, bright and undergoes rapid and complete photoconversion. We have compared PA-GFP, PS-CFP2, Kaede and KikGR four readily available and commonly used photomodulatable fluorescent proteins for use in ES cells and mice. Photomodulatable fluorescent proteins are one of the emerging classes of genetically-encoded fluorescent proteins. With the advent of more accessible and sophisticated imaging technologies as well as an abundance of fluorescent proteins with different spectral characteristics, the dynamic processes taking place in situ in living cells and tissues can now be probed. The use of genetically-encoded fluorescent proteins has revolutionized the fields of cell and developmental biology and in doing so redefined our understanding of the dynamic morphogenetic processes that shape the embryo.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
